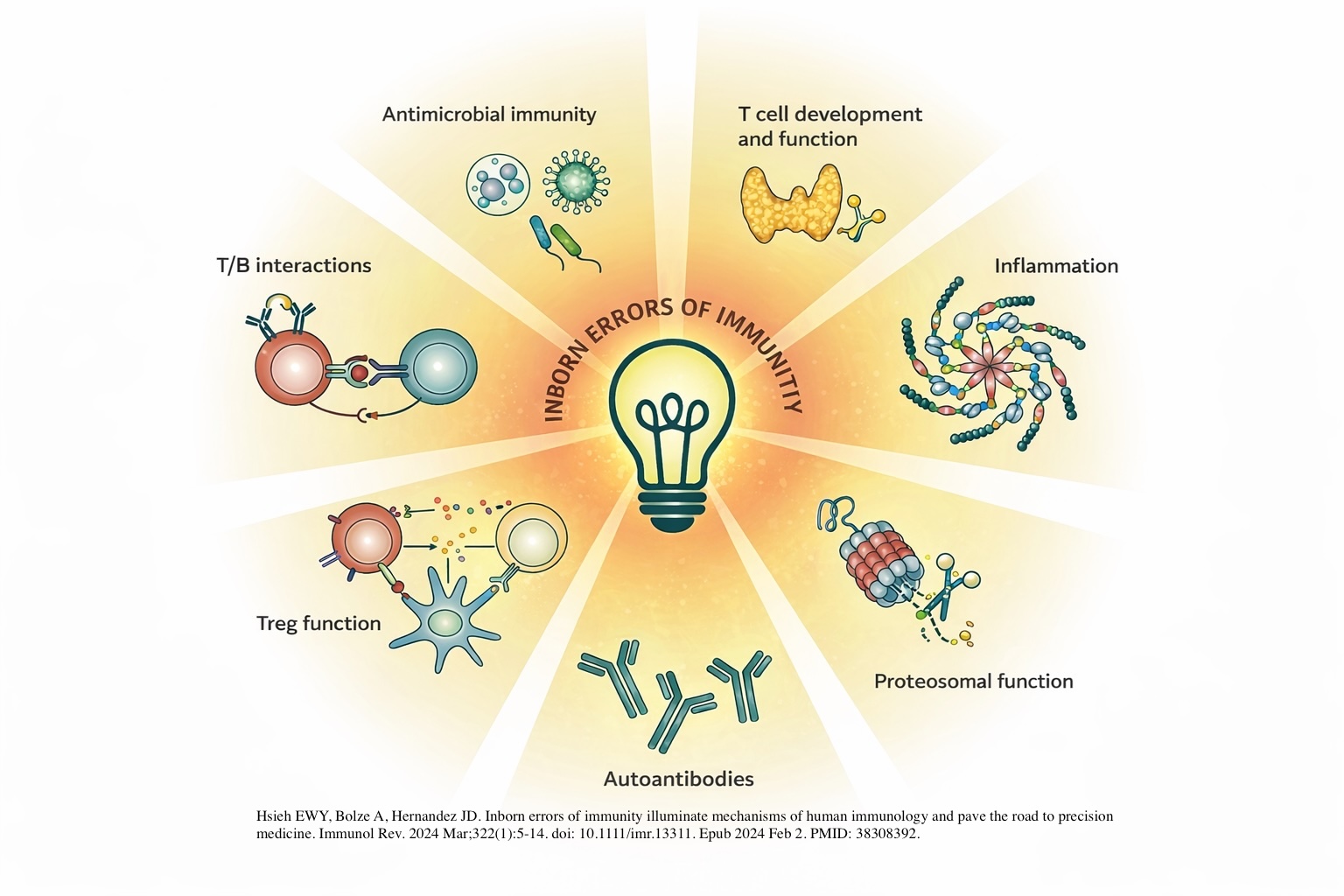
Exploration of Novel IEIs
Understanding why only a small subset of individuals develop life-threatening infections, immune dysregulation, or malignancy reveals fundamental principles and non-redundant immune pathways governing disease pathogenicity in humans.
Background
Interindividual variability in susceptibility to infection, immune dysregulation, and malignancy risk remains a fundamental and largely unanswered question in human biology. Even among individuals exposed to the same pathogens, outcomes range from asymptomatic infection to life-threatening disease, highlighting critical gaps in our understanding of protective and pathological immunity.
-
Extreme clinical variability during infection and immune dysregulation reflects underlying genetic and immunological determinants that are poorly understood
(Annu Rev Pathol, 2024; PNAS, 2015). -
Despite major advances in sequencing technologies, the diagnostic yield for patients with suspected inborn errors of immunity remains limited, at approximately 30%
(Frontiers in Pediatrics, 2024), suggesting that many disease-causing genes, allelic variants, and mechanisms remain undiscovered. -
Identifying and characterizing novel IEIs not only enables precise diagnosis for the patients we study, but also uncovers essential immune pathways that govern host defense, immune regulation, and malignancy risk in humans.
Key Questions We Ask
- Why do some individuals develop severe or life-threatening disease after otherwise common infections?
- Which immune pathways are non-redundant in humans causing or preventing patients from diseases of interests?
- How do monogenic defects predispose to immune dysregulation and cancer?
Disease Phenotypes We Study
- Life-threatening infections, with a particular focus on Epstein–Barr virus (EBV)
- Immune dysregulation and malignancy in the context of EBV infection
- Broader spectra of immune dysregulation, including autoimmunity and autoinflammation
Our Approach
Our work is grounded in patient-oriented research and integrates:
- Patient-driven discovery, including deep clinical phenotyping, enrollment, and next-generation sequencing
- Human genetics and immunology, to link genotype to cellular and molecular immune phenotypes
- Deep mechanistic investigation, to establish causal relationships between genetic defects and disease
Rather than beginning with predefined pathways, we prioritize unbiased discovery in humans, followed by rigorous functional validation in relevant immune systems.
Selected Publications
(As we are in the process of building our independent research program, representative publications are drawn from the PI’s prior training.)
Yang R, Mele F, Worley L, Langlais D, Rosain J, Benhsaien I, et al.
Human T-bet governs innate and innate-like adaptive IFN-γ immunity against mycobacteria.
Cell. 2020;183(7):1826–1847.e31.
DOI: 10.1016/j.cell.2020.10.046
Yang R, Avery DT, Jackson KJL, Ogishi M, Benhsaien I, Du L, et al.
Human T-bet governs the generation of a distinct subset of CD11c^high^ CD21^low^ B cells.
Science Immunology. 2022;7(73):eabq3277.
DOI: 10.1126/sciimmunol.abq3277
Yang R, Weisshaar M, Mele F, Benhsaien I, Dorgham K, Han J, et al.
High Th2 cytokine levels and upper airway inflammation in human inherited T-bet deficiency.
Journal of Experimental Medicine. 2021;218(8):e20202726.
DOI: 10.1084/jem.20202726